27-Apr-2026
Predicting metal binding sites is key for understanding their function and has a wide variety of applications, including designing metal-targeting therapeutics against viruses such as SARS-CoV-2 (the virus which causes COVID-19). There are several existing tools which can predict metal binding sites within a protein based on its structure; however, these tools often rely on comparisons to existing protein structures to make predictions. The team of Antonio Rosato, based at the University of Florence in Italy, recently developed a new prediction model which can predict zinc metal ion binding sites without dependence on existing protein structures.
The team included Vincenzo Laveglia (first author on the paper), Cosimo Ciofalo, Enrico Morelli, and Claudia Andreini. They developed the “Master of Metals 2” model through machine learning approaches, where an algorithm learns to perform a task by being given some training data. In the case of Master of Metals 2 (MoM2), the training data consisted of a curated dataset taken from the Protein Data Bank (PDB), consisting of 1294 structures of zinc-bound proteins with 1803 target sites.
The prediction of MoM2 can be broken down into three steps. First, amino acids from the protein of interest are classified as being either part of the metal binding site, next to an amino acid which is part of the binding site, or not part of the metal binding site at all. This decision is based on the properties of the graph representing the protein structure around each amino acid (shown in Figure 1). Secondly, once all metal binding site amino acids have been assigned, these are then grouped and assigned to a particular binding site. This step uses the BioMetAll algorithm originally developed by the group of Jean-Didier Maréchal from the Autonomous University of Barcelona. Essentially, animo acids are assigned to binding sites using distance and angle constraints extracted from existing scientific literature. In the final stage, the predictions made by MoM2 are assessed using a set of feed forward neural networks. Each site is given a score, and if amino acids are predicted to belong to multiple sites, then the site with the highest score is chosen, to reduce redundancy.
After training a model, the final step is to test how well its predictions work using an independent data set. MoM2 is a type of machine learning model which performs a “binary task”; one where the answer is yes or no. This type of model can be assessed using three statistics: precision, recall (also known as sensitivity), and the F1 score. Precision essentially asks, “of all the metal binding sites we predicted, how many were true metal binding sites, and how many were false positives?”. Recall asks, “of all the true metal binding sites that existed, how many did we detect?”. Generally, a model can be optimised to increase precision at the expense of recall, and vice-versa, and so the F1 score considers both the precision and the recall.
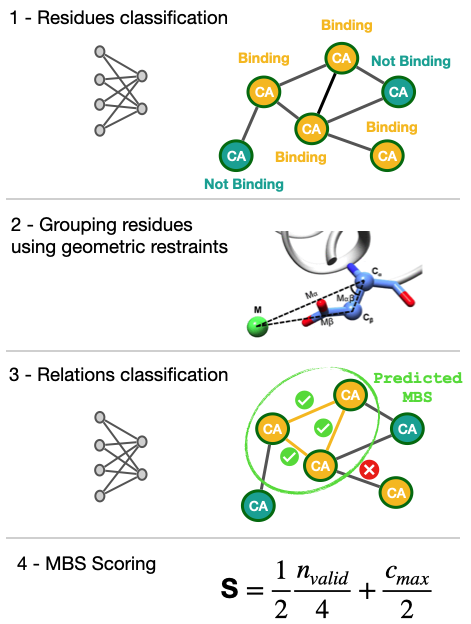
Figure 1 – Overview of how Master of Metals2 (MoM2) predicts zinc metal ion binding sites.
To test whether it was an improvement over existing tools, MoM2 was benchmarked using an established procedure. This test involved using comparable tools to predict metal binding sites for proteins not included in the training data. These proteins already had an experimentally determined structure, against which predictions could be verified. MoM2 outperformed every other prediction tool that was tested, demonstrating greater precision, recall, a higher F1 score, and a lower rate of false positives. Importantly, while MoM2 can be optimised for precision at the expense of recall (and vice versa), optimising in one direction did not lead to a large decrease in performance in the other direction.
As mentioned above, determining metal binding sites could have implications for anti-viral therapies which target the essential metals within viral proteins. Therefore, MoM2 was used to predict zinc metal ion binding sites in SARS-CoV-2, and the results compared to previously predicted sites. MoM2 correctly predicted 18 of the expected 20 binding sites within the SARS-CoV-2 proteins and produced only 2 false positives. This is particularly impressive given that the dataset on which MoM2 was trained contained only 2 SARS-CoV-2 proteins.
Besides the lack of reliance on similarity to existing structures to make predictions, and the improved performance, MoM2 also represents another great improvement over existing tools: accessibility. MoM2 is freely available online using a web portal, and so does not require the user to download any software or have knowledge of programming languages like Python. MoM2 accepts inputs in the form of protein structures, PDB IDs, or UniProt IDs, all of which will be familiar to members of the structural biology community. MoM2 also works quickly - a prediction for a whole bacterial proteome can be obtained from the web server in a few hours.
“ITACA.SB funding has been crucial to secure the GPU infrastructure as well the other hardware and software that we have used for the development and implementation of MoM2. We believe that is will be a very useful tool to the community bioinorganic chemists who are interested in the identification of metal binding sites as part of their research to understand the biological role of metals (zinc in this specific case).”
Says Antonio Rosato, Associate Professor at the University of Florence and leader of the research group behind MoM2.
You can the read the full paper, published in Briefings in Bioinformatics here - “Master of Metals2: a graph neural network based architecture for the prediction of zinc binding sites in protein structures”. The work was supported by funding from several projects, including Potentiating the Italian Capacity for Structural Biology Services in Instruct-ERIC ("ITACA.SB"), Fragment-Screen, and W-BioCat. If you have used Instruct-ERIC infrastructure to collect data for a manuscript and would like to have it published as a Science Highlight, then please get in touch with us!