17-Mar-2026
EMBL-EBI have launched a new collaboration with NVIDIA, Google DeepMind, and Seoul National University, which aims to make millions of AI predictions for protein complex structures openly available to the scientific community through the AlphaFold Database.
This initial data release is the next step in an ambition to double the size of the AlphaFold Database, co-developed by EMBL-EBI and Google DeepMind in 2021, giving scientists worldwide access to high-confidence AI predictions for protein complexes.
Spanning the most important protein complexes for studying human health and disease, this is the largest dataset of its kind to date, prioritising proteins from the most studied 20 organisms, including human, and the World Health Organization’s bacterial priority pathogens list.
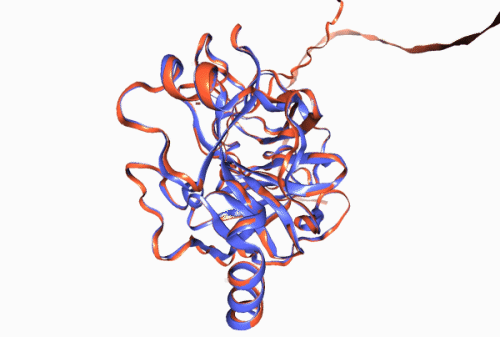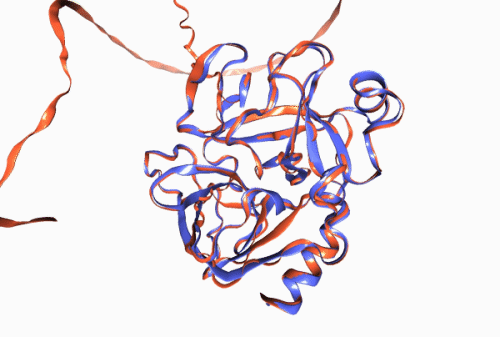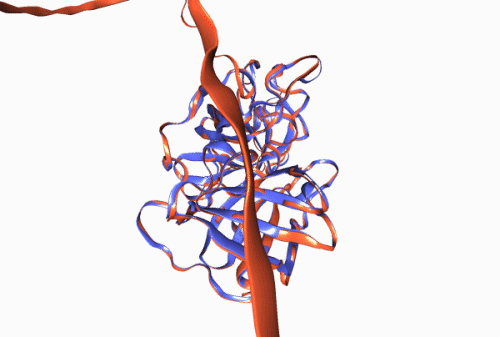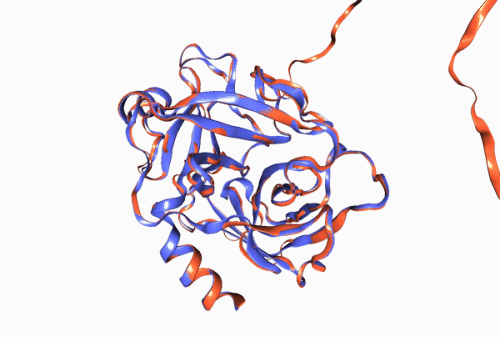
Figure 1. Experimentally derived (blue) and AlphaFold prediction (red) of Trypsin 3 protein structure.
EMBL-EBI convened the collaboration, and contributed deep biological knowledge, as well as expertise in biodata management and analysis. The Steinegger Lab at Seoul National University developed the methodology, based on Google DeepMind’s AI system AlphaFold. NVIDIA provided cutting-edge accelerated compute infrastructure and improvements to data pipelines to overcome limitations that historically made this scale of calculations challenging. Together with Google DeepMind, EMBL-EBI integrated the new dataset into the AlphaFold Database.
Instruct Director Prof. Harald Schwalbe: “As structural biologists, we are happy to see that the experimental data serve for these new initiatives. We are happy that EMBL-EBI, a partner in Instruct-ERIC, plays a key role in this. We believe that mining the richness of experimental raw data in the future will pave the way for improved AI prediction. The breadth of structural biology data and their integration in integrated scientific biology will be essential. Instruct-ERIC is playing a key role in making such services and exploitation of results available implementing FAIR principles.”
AlphaFold has already calculated predictions for 30 million complexes. Of these, 1.7 million high-confidence homodimer predictions have been added to the AlphaFold Database. Another 18 million are lower-confidence homodimers, which will be made available as a list and for bulk download from the EMBL-EBI FTP server in the coming days.
Find out more about AI and structural biology, as well as a range of computational tools available through Instruct-ERIC here.
